


Next: 11 Unsampled subpopulations
Up: Inferring patterns of migration
Previous: 9 Plotting predicted and
Contents
The model is able to handle most forms of sampling error.
Typically, only a proportion of individuals are sampled in
each subpopulation. The most realistic assumption is to
assume that the number of copies of, say allele 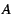 ,
conditional on the allele frequency
,
conditional on the allele frequency 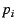 in subpopulation
in subpopulation 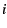 follow a hypergeometric distribution. If we have
hypergeometric sampling of
follow a hypergeometric distribution. If we have
hypergeometric sampling of 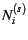 individuals from a
finite population of
individuals from a
finite population of 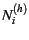 individuals, then the
conditional variance in the sampled gene frequency is
individuals, then the
conditional variance in the sampled gene frequency is
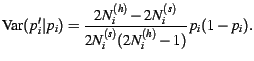 |
(5) |
The unconditional standardized variance of the sampled gene
frequency is then
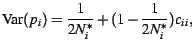 |
(6) |
where
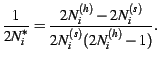 |
(7) |
This simplifies to the more standard binomial model when
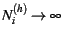 . Also, as
. Also, as
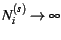 there sampling variance tends to zero.
there sampling variance tends to zero.
To set up the model with sampling error, the two vectors
Ns and Nh containing the appropriate sample
sizes in each subpopulation (corresponding to 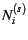 and
and 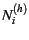 , respectively) should be defined globally.
The diagonal elements of the computed covariance matrix are
then adjusted according to (6), before evaluating
the likelihood of the data. In addition, in the simulate procedure, sampling error is added at the end of
each simulated realization of the process. Sample sizes
differing between loci is currently not handled. Assigning
the value Inf to all elements of Nh results in
binomial sampling. The model is set up with no sampling
error by assigning a NA to Ns or by assigning
Inf to some (or all) of the elements.
, respectively) should be defined globally.
The diagonal elements of the computed covariance matrix are
then adjusted according to (6), before evaluating
the likelihood of the data. In addition, in the simulate procedure, sampling error is added at the end of
each simulated realization of the process. Sample sizes
differing between loci is currently not handled. Assigning
the value Inf to all elements of Nh results in
binomial sampling. The model is set up with no sampling
error by assigning a NA to Ns or by assigning
Inf to some (or all) of the elements.



Next: 11 Unsampled subpopulations
Up: Inferring patterns of migration
Previous: 9 Plotting predicted and
Contents
Jarle Tufto
2001-08-28
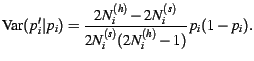
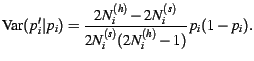
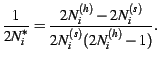
![]() and
and ![]() , respectively) should be defined globally.
The diagonal elements of the computed covariance matrix are
then adjusted according to (6), before evaluating
the likelihood of the data. In addition, in the simulate procedure, sampling error is added at the end of
each simulated realization of the process. Sample sizes
differing between loci is currently not handled. Assigning
the value Inf to all elements of Nh results in
binomial sampling. The model is set up with no sampling
error by assigning a NA to Ns or by assigning
Inf to some (or all) of the elements.
, respectively) should be defined globally.
The diagonal elements of the computed covariance matrix are
then adjusted according to (6), before evaluating
the likelihood of the data. In addition, in the simulate procedure, sampling error is added at the end of
each simulated realization of the process. Sample sizes
differing between loci is currently not handled. Assigning
the value Inf to all elements of Nh results in
binomial sampling. The model is set up with no sampling
error by assigning a NA to Ns or by assigning
Inf to some (or all) of the elements.